Analysis of U-Net Neural Network Training Parameters for Tomographic Images Segmentation
Latin-American Journal of Computing, vol. 10, no. 2, pp. 84-95, 2023
Escuela Politécnica Nacional
Received: 13 March 2023
Accepted: 17 May 2023
Abstract: In the field of computational vision, image segmentation is one of the most important resources. Nowadays, this procedure can be made with high precision using Deep Learning, and this fact is important to applications of several research areas including medical image analysis. Image segmentation is currently applied to find tumors, bone defects and other elements that are crucial to achieve accurate diagnoses. The objective of the present work is to verify the influence of parameters variation on U-Net, a Deep Convolutional Neural Network with Deep Learning for biomedical image segmentation. The dataset was obtained from Kaggle website (www.kaggle.com) and contains 267 volumes of lung computed tomography scans, which are composed of the 2D images and their respective masks (ground truth). The dataset was subdivided in 80% of the volumes for training and 20% for testing. The results were evaluated using the Dice Similarity Coefficient as metric and the value 84% was the mean obtained for the testing set, applying the best parameters considered.
Keywords: Deep Learning, Biomedical Image Segmentation, Fully Convolutional Networks, U-Net, Computed Tomography.
I. Introduction
Deep Learning (DL) is a branch of machine learning developed by learning successive layers, almost always using models called neural networks [1], through data representations. Nowadays the application of DL has presented promising results in biomedical image segmentation. Zhang et al. [2] designed Convolutional Neural Networks (CNNs) [3] architectures to segment infant brain tissues in Magnetic Resonance (MR) images which is a process even more difficult for adults due to the low tissue contrast, increased noise and ongoing white matter myelination. The CNNs segmentation of isointense-phase brain image outperformed competing methods on a set of manual process. Oktay et al. [4] applied the U-Net model to the segmentation of pancreas, which presented Dice Similarity Coefficients (DSCs) 2% to 3% higher than other models. This improvement in pancreas segmentation is important in many clinical applications of liver segmentation in 3D images [5]. In addition, through the application of DL, it becomes possible to perform more qualitative or even quantitative analysis of the regions of interest, such as lesions [6], a factor that represents important advances for the entire healthcare sector. The use of DL generates results that do not rely on the subjectivity of the observer and provide a decrease of the time needed for segmentation once the model is trained.
Although several Artificial Neural Networks (ANNs) [7] have been used in biomedical image segmentation, in this work, the U-Net architecture was used. U-Net is a Fully Convolutional Neural Network (FCN or Fully Convnet) that has been consolidating as one of the most prominent ANNs for biomedical images.
The performance improvement of U-Net into images segmentation can be achieved by adjusting the parameters for the problem in question. Goyal et al. [8] investigated the influence of batch size in other CNNs performances and verified that large minibatches cause optimization difficulties when working with ImageNet dataset [9]. Furthermore, Nishio et al. [10] developed a methodology for determining cancer diagnoses, classifying nodules between benign and malignant as well as verifying the stage of the disease, which could correspond to primary lung cancer or metastatic lung cancer. Evaluating the effect of image size as input of the deep convolutional neural network used, testing image sizes equal to 56, 112 and 224, the results showed that larger image sizes as inputs improved the accuracy of lung nodule classification.
The objective of the present work is to evaluate the influence of U-Net parameters variation during the segmentation of lung computed tomographies. This way, it is possible to segment the lungs in the tomography with high precision which makes subsequent processes easier, for example, the detection of pulmonary nodules for early diagnosis of lung cancer.
The remaining of the article is structured as follows. Section II contains the theoretical framework. Section III presents the adopted methodology. Section IV is dedicated to the results and discussions. Finally, section V contains the conclusion of the work.
II. Theorical Framework
A. Biomedical Image Segmentation
Image segmentation is considered the most important medical imaging process and corresponds to the extraction of the Region of Interest (ROI), subdividing the image into areas based on specific characteristics, such as segmenting body organs and tissues to detect tumors and other elements and this division can be applied for both 2D and 3D data [11]. The binary segmentation, applied in the present work, subdivides the image in a white part that corresponds to the mask containing the ROI, and a black part as background. Once obtained, in addition to detecting tumors or other abnormalities, the masks can measure tissue growth analyzing the growth of possible tumors and help in treatment planning.
B. Fully Convolutional Neural Networks
FCNs are classified as a specific type of CNNs that contains a convolutional path connected to a Fully Connected (FC) layer [12]. The FCs has Multilayer Perceptron (MLP) as main representative and they are used to classification. The main difference between the designs of CNNs and FCNs is that the last one has a deconvolutional path, also called expansion path, instead of the dense layer. Therefore, FC reduce the number of parameters once there are no FC layers, speeding up learning and inference. As output, the FCNs generate a pixel vector whose size corresponds to the input data [13].
C. Net
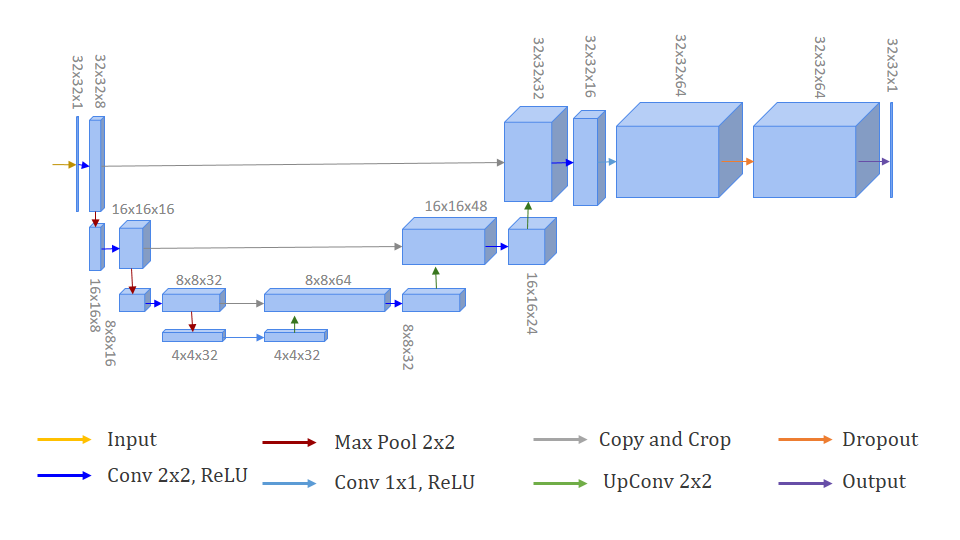
U-Net is an FCN created by Ronneberger, Fischer and Brox [14] in order to cope with biomedical images segmentation. The architecture of the ANN is composed of two paths. The first path is called the contracting one, which aims to capture the context. On the other hand, the second path corresponds to the expanding path, which allows precise localization. Precisely, because of the fact that the database of biomedical images uses not to be very large, the authors created a network that is able to be trained end-to-end and presents accurate results.
In Fig. 1, each blue box corresponds to a multi-channel feature map. The path indicated by the red arrows on the left side of the image corresponds to convolutional path, also known as downsampling path. On the right side, indicated by green arrows, there is the deconvolutional path or upsampling path. Just before the output, there is a layer called Dropout, where some neurons are randomly turned off during the training. This regularization is a method to avoid overfitting in the process.
D. Batch Size
During each epoch, all training set was used but subdivided in batches, for updating the U-Net weights and improving performance. Batch Size is the hyperparameter in Convolutional Neural Networks that corresponds to the number of images used to train a single forward and backward pass. The correlation between CNNs performance and Batch Size also depends on the datasets nature, mainly in the case of medical ones due to its complexity [15].
E. Early Stopping
Early Stopping is considered a regularization method capable of determining when to stop the execution of an iterative algorithm [16]. This callback calculates the precision of segmentation or classification using validation data. It interrupts the training when precision stops improving, avoiding overfitting, within a given range called patience that corresponds to its most important hyperparameter.
F. Related Works
Paiva et al. [17] used U-Net to segment microcomputed tomography images which the ROI corresponds to lenses of tadpole specimen of the frog Thoropa miliaris and compared the performance to methods of semiautomatic segmentation. The research concluded that the automatic segmentation using Fully Convnet was much faster than the semiautomatic processes and it also showed high accuracy.
Moura and Meneses [18] segmented heart computed tomography images testing U-Net performance with number of epochs (50, 100) variation, number of features (32, 64) variation, BatchNormalization, RMSprop optimizer function and BinaryCrossentropy loss function. The authors concluded that there was no statistically significant difference between the different parameters adjustment, therefore the chosen model could be the one with the smallest standard deviation (0.065) and the smallest time of execution (368 seconds), which was U-Net with 32 feature maps, BatchNormalization and 100 epochs.
Saood and Hatem [19] segmented lung Computed Tomography (CT) using U-Net and SegNet in order to detect and label infected tissues in the lungs and contribute to verify the diagnosis of patients contaminated by COVID-19. The results of the work presented that U-Net showed better performance as a multi-class segmentor.
Kandel and Castelli [15] classified images from a histopathology dataset, testing batch size equal to 16, 32, 64, 128 and 256, with fixed learning rate 0.001 and Adam optimizer. The authors concluded that the largest batch size presented the highest performance.
Thambawita et al [20] classified gastrointestinal endoscopy images using two different CNNs models testing image resolutions ranging from 32x32 to 512x512 pixels. The results showed that the best performance occurred when the models were training and applied into testing data with the highest image resolution.
III. Material and Methods
The code was implemented in Python 3.8.13, using Keras 2.6.0 API, Numpy 1.21.5, and OpenCV 4.5.5.64 with Tensorflow 2.6.0 as backend. An HP Elitedesk 800 desktop was used with an Intel Core i7-6700 3.40 GHz CPU, 16 GB RAM, Windows 10 Pro 64 bits Operating System, and an Nvidia GeForce GT 730 GPU.
Fig. 2 presents the research methodological flowchart and each stage will be described during the following sections.

A. Dataset
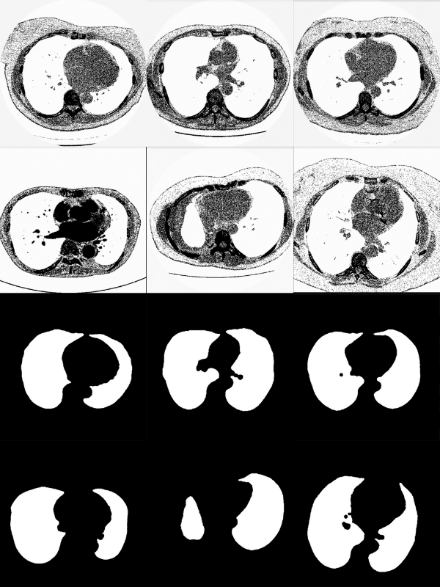
The CT volumes were downloaded from Kaggle website (https://www.kaggle.com/datasets/kmader/finding-lungs-in ct-data?resource=download) which contains several open source datasets.
The database used is composed of 267 lungs CT scans and each volume presents 2D images and their corresponding mask representing the ROI highlighted in white and the background in black. The data contained 512x512 pixels on its dimensions.
B. Data Preprocessing
Before starting the segmentation, there were three stages of preprocessing that had to be performed. First step was to subdivide the volumes in training and testing. Therefore, 80% of the initial data became training and validation data, corresponding to 214 volumes, while 20% of the initial data became testing data, equivalent to 53 volumes. Secondly, the images were resized for ensuring the correct reading of the data and avoiding possible differences in file dimensions.
Thirdly, the images were normalized. Let I be an n-dimensional grayscale image. A linear normalization transforms 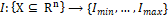 with intensity values in the range
with intensity values in the range  , into a new image
, into a new image  with intensity values in the range
with intensity values in the range  . The linear normalization is represented by:
. The linear normalization is represented by:
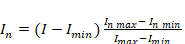 (1)
(1)where 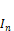 is the new intensity,
is the new intensity,  is the initial intensity,
is the initial intensity,  is the desired minimum intensity, is the desired maximum intensity
is the desired minimum intensity, is the desired maximum intensity  and
and 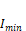 and are the current maximum and minimum
intensity. In this case, the minimum
value of the range is equal to 0 while the maximum one is equivalent to 1.
and are the current maximum and minimum
intensity. In this case, the minimum
value of the range is equal to 0 while the maximum one is equivalent to 1.
C. Data Augmentation
Once the preprocessing was completed, a process of Data Augmentation [21] was realized to enable an increase in the number of training volumes based on the original ones. In order to add more variability in the dataset, certain geometric transformations were applied, such as horizontal and vertical displacement of the lungs in the images, rotation and zoom. With these changes, from each original image 32 new images were generated, improving quality of the training data and avoiding overfitting.
D. Callbacks
Besides Early Stopping as mentioned in section II, the model was performed using two other callbacks, Model Checkpoint and Learning Rate Scheduler, respectively to save the best weight configuration in a .h5 format file and to update the value of the optimization rate between the epochs.
E. Data Splitting
Furthermore, of the 214 volumes reserved for training, only 80% (171 volumes) were actually used for training while the remaining 20% (43 volumes) were used for model validation.
F. Dice Similarity Coefficient
The metric used for both training and testing was the DSC. The DSC corresponds to a comparison between the result shown by the model and the mask equivalent to the image segmented that is the Ground Truth (GT), the reference image. The metric is a reason between the double of the intersection of the compared images and the sum of pixels of the both images and can be calculated by the equation below:
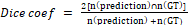 (2)
(2)G. Parameter testing
Different numbers of epochs and patience value from Early Stopping were preliminarily tested. The number of epochs tested were 10, 50 and 100. Then, patience values tested of Early Stopping were 10, 20, 30, 40 and 50. Subsequently, Batch Size values equal to 4, 8, 16 and 32 were tested. Finally, Image Size equal to 32, 64 and 128 pixels were implemented. Five executions were performed for each parameter test in order to analyze its influence on model performance. The seed used during each execution was randomized with a range of 1 to 100 in order to guarantee impartiality in the process.
H. Statistical Analysis
Kruskal-Wallis and Dunn’s tests [22, 23] were performed in order to verify if there is statistically significant difference between the parameters’ values tested. The Kruskal-Wallis test is used to compare three or more groups of data. If the null hypothesis of no statistically significant difference is rejected, then the Dunn’s post-hoc test is used for pairwise comparison between the groups. The Bonferroni correction was applied to Dunn’s test for reducing the Type I Error probability. The threshold 0.05 was adopted for the statistical tests. The graphs were plotted with Plotly 5.11.0.
IV. Results and Discussion
A. Preliminary tests
Kruskal-Wallis test was implemented to analyze the results of number of epochs preliminary test and the p-value obtained was equal to 0.6861. The value shows that there is no statistically significant difference in model performance using different number of epochs, but once using 50 epochs the segmentation presented the highest mean (0.7873) and also the smallest standard deviation (0.0153), it was selected to next steps.
The same procedure was applied to evaluate the results of Early Stopping patience value preliminary test. The p-value was equal to 0.4305, which points to the fact that also no statistically significant difference was found. Again, once patience value equal to 40 obtained the highest mean (0.7880) with the smallest standard deviation (0.0170), it was chosen as the best parameter to patience.
B. Batch Size
In Fig. 4, it is possible to verify the boxplots of DSCs distribution for Batch Size comparison. Table I presents the statistical results for Kruskal-Wallis test.

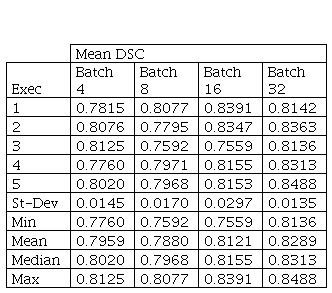
In this case, the p-value obtained with Kruskal-Wallis test was equal to 0.028 between batch size values used. Then, Dunn's multiple comparisons test with Bonferroni correction was applied to the results.
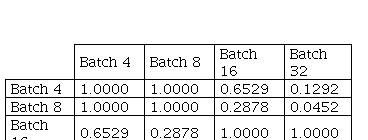
Although there is no statistically significant difference between the results obtained using batch size 16 and 32, batch size 32 was statistically significant different from batch size 8, with p-value 0.0452. Batch size 32 also presented a tendency for better results, with mean 0.8289 and standard deviation 0.0135.
C. Image Size
In Fig. 5, it is possible to verify the boxplots of DSCs distribution for Image Size comparison. Table III presents the statistical results for Image Size parameter.

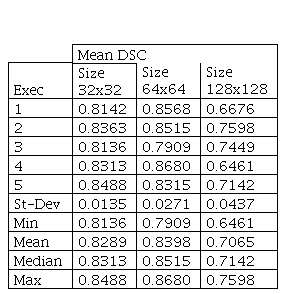
The p-value obtained by Kruskal-Wallis was 0.0068 between image sizes used. Therefore, Dunn’s test with Bonferroni correction was also implemented and the results are presented as follows.
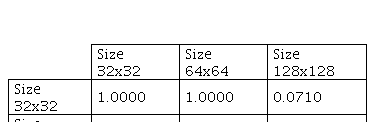
he Dunn’s test results show that there is no statistically significant difference between the means obtained using image size equal to 32 and 64. There is only a statistically significant difference between 64×64 with respect to 128×128. Furthermore, although image size 32 presented the smallest standard deviation (0.0135), once image size 64 showed the highest mean (0.8398) and the highest median (0.8515), it was chosen as the best parameter.
Fig. 6 presents the segmentation with maximum DSC. It is possible to verify that the lung in the tomography has well-defined edges and low noise.

Conversely, Fig. 7 shows the segmentation with minimum DSC, showing that the CT scan has high noise and low contrast on its edges.

This way, it is observed that the best segmentations occurred to images that had low noise image, the lung borders are not scattered and there was a significant contrast between the lung and background.
V. Conclusion
In this work, U-Net was used to segment lungs in CT data. The dataset was subdivided in training data and testing data and the model was trained using the masks (GTs) also available in the dataset.
Four parameter tests were implemented. First, different number of epochs and early stopping patience were compared with no statistically significant difference observed.
Afterwards, Batch Size test was performed, in which 32 was classified as the most satisfactory value, demonstrating that the model needed to increase the number of images used to training on each forward and backward pass, probably because of computed tomography complexity.
Finally, Image Size test was performed, in which 64×64 pixels was the dimension that presented better results in relation to 128×128, it shows that using the highest dimensions available can cause difficulty to model learning and segmentation, but the results were not conclusive regarding 32×32 pixels.
Predictions performed by the model to the testing data were compared to the GT, presenting satisfactory results in terms of the metrics used as basis throughout the process (DSC) even though the low number of volumes in the dataset.
Therefore, in terms of the dataset used, the present research confirms the efficiency of the U-Net architecture in order to segment biomedical images, factor which enables its implementation to future works of interest to both DL and health care applications.
Acknowledgment
A.A.M.M., R.C.B. and Y.S.P. acknowledge CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico). R.C.B. thanks FAPERJ (Fundação de Amparo à Pesquisa do Rio de Janeiro). D.G.S. thanks CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior).
References
[1] F. Chollet, Deep Learning with Python. Manning Publications; 2017. ISBN 9781617294433.
[2] W. Zhang et al., “Deep convolutional neural networks for multi-modality isointense infant brain image segmentation,” Neuroimage, vol. 108, pp. 214–224, Mar. 2015, doi: 10.1016/j.neuroimage.2014.12.061.
[3] Y. Lecun, E. Bottou, Y. Bengio, and P. Haffner, “Gradient-Based Learning Applied to Document Recognition”, 1998.
[4] O. Oktay et al., “Attention U-Net: Learning Where to Look for the Pancreas,” Apr. 2018, [Online]. Available: http://arxiv.org/abs/1804.03999
[5] D. Shen, G. Wu, and H.-I. Suk, “Deep Learning in Medical Image Analysis,” 2017, doi: 10.1146/annurev-bioeng-071516.
[6] X. Liu, L. Song, S. Liu, and Y. Zhang, “A review of deep-learning-based medical image segmentation methods,” Sustainability (Switzerland), vol. 13, no. 3, pp. 1–29, Feb. 2021, doi: 10.3390/su13031224.
[7] S. Haykin, Neural Networks: A Comprehensive Foundation., Prentice-Hall, 1999.
[8] P. Goyal et al., “Accurate, Large Minibatch SGD: Training ImageNet in 1 Hour,” Jun. 2017, [Online]. Available: http://arxiv.org/abs/1706.02677.
[9] J. Deng, W. Dong, R. Socher, L.-J. Li, K. Li, and L. Fei-Fei, ImageNet: A Large-Scale Hierarchical Image Database. IEEE, 2009.
[10] M. Nishio et al., “Computer-aided diagnosis of lung nodule classification between benign nodule, primary lung cancer, and metastatic lung cancer at different image size using deep convolutional neural network with transfer learning,” PLoS One, vol. 13, no. 7, Jul. 2018, doi: 10.1371/journal.pone.0200721.
[11] A. Ashour, Y. Guo, and Mohamed W., “Medical Image Segmentation.”
[12] I. Goodfellow, A. Courville and Y. Bengio. Deep Learning (Adaptive Computation and Machine Learning Series).
[13] D. Nie, L. Wang, Y. Gao, and D. Sken, “Fully convolutional networks for multi-modality isointense infant brain image segmentation,” in Proceedings - International Symposium on Biomedical Imaging, Jun. 2016, vol. 2016-June, pp. 1342–1345. doi: 10.1109/ISBI.2016.7493515.
[14] O. Ronneberger, P. Fischer, and T. Brox, “U-net: Convolutional networks for biomedical image segmentation,” in Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 2015, vol. 9351, pp. 234–241. doi: 10.1007/978-3-319-24574-4_28.
[15] I. Kandel and M. Castelli, “The effect of batch size on the generalizability of the convolutional neural networks on a histopathology dataset,” ICT Express, vol. 6, no. 4, pp. 312–315, Dec. 2020, doi: 10.1016/j.icte.2020.04.010.
[16] G. Raskutti, M. J. Wainwright, and B. Yu, “Early Stopping and Non-parametric Regression: An Optimal Data-dependent Stopping Rule,” 2014.
[17] K. Paiva et al., “Performance evaluation of segmentation methods for assessing the lens of the frog Thoropa miliaris from synchrotron-based phase-contrast micro-CT images,” Physica Medica, vol. 94, pp. 43–52, Feb. 2022, doi: 10.1016/j.ejmp.2021.12.013.
[18] M. Moura and A. Meneses, “Evaluation Of Unet Convolutional Neural Network Parameters For Segmentation Of Heart CT Images”. Available in the annals of XXIV National Meeting of Computacional Modeling (ENMC).
[19] A. Saood and I. Hatem, “COVID-19 lung CT image segmentation using deep learning methods: U-Net versus SegNet,” BMC Med Imaging, vol. 21, no. 1, Dec. 2021, doi: 10.1186/s12880-020-00529-5.
[20] V. Thambawita, I. Strümke, S. A. Hicks, P. Halvorsen, S. Parasa, and M. A. Riegler, “Impact of image resolution on deep learning performance in endoscopy image classification: An experimental study using a large dataset of endoscopic images,” Diagnostics, vol. 11, no. 12, Dec. 2021, doi: 10.3390/diagnostics11122183.
[21] C. Shorten and T. M. Khoshgoftaar, “A survey on Image Data Augmentation for Deep Learning,” J Big Data, vol. 6, no. 1, Dec. 2019, doi: 10.1186/s40537-019-0197-0.
[22] A. Dmitrienko, C. Chuang-Stein, R. D’Agostino. “Pharmaceutical Statistics Using SAS”, 2014. In Journal of Chemical Information and Modeling (Vol. 53, Issue 9). SAS Publishing.
[23] M. Neuhauser. “Nonparametric Statistical Tests”, 2011. Chapman and Hall/CRC. https://doi.org/10.1201/b11427.

