1. Introduction
In recent years, cannabis cultivation (Cannabis sativa L.) has spread throughout the vast majority of Uruguayan departments, with a greater influence in the south and east of the country(1).
Being a relatively new crop in the country, it is important to know the health problems that can affect it. In this regard, pests and fungi associated with diseases in five cannabis crops were inspected during the production cycle in 2018 and 2019, in the south of the country. A total of 11 plants with symptoms of stem canker were observed in three inflorescence-production crops, from which Neofusicoccum sp.(2) was isolated.
Neofusicoccum parvum ((Pennycook & Samuels) Crous, Slippers & A.J.L. Phillips 2006) along with other species in the Botryosphaeriaceae family cause canker and blight in several crops and woody species(3). Moreover, the pathogen was first reported in cannabis crops in Italy(4) and subsequently in the United States(5). In Uruguay, N. parvum affects the crops of apple tree(6)(7), pear, peach(6) and vine(8), but it is unknown whether it can cause the disease in cannabis. Thus, this study aims to characterize three isolates of Neofusicoccum by morphological and molecular analysis, and pathogenicity tests in cannabis plants.
2. Material and methods
2.1 Isolates of Neofusicoccum
Three of 11 purified isolates of Neofusicoccum sp. were used from the Phytopathology Laboratory collection (Agronomy College, Udelar). The isolates were obtained from plants with canker symptoms in stems collected during 2018 and 2019 (Table 1).
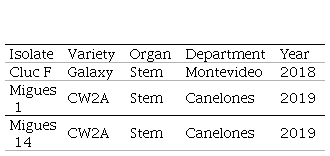
Table 1
Isolates of Neofusicoccum sp.
|
Isolate
|
Variety
|
Organ
|
Department
|
Year
|
|
Cluc F
|
Galaxy
|
Stem
|
Montevideo
|
2018
|
|
Migues 1
|
CW2A
|
Stem
|
Canelones
|
2019
|
|
Migues 14
|
CW2A
|
Stem
|
Canelones
|
2019
|
2.2 Morphological identification
Each isolate was placed onto Petri dishes for morphological identification, with potato dextrose agar (PDA) medium (Oxoid Ltd., Hampshire, England) at 25 °C, and colony characteristics were observed at 3, 8 and 16 d incubation, using the description by Pennycook & Samuels(9) and Phillips and others(10). Furthermore, the isolates were incubated on water agar (WA) medium (Oxoid Ltd., Hampshire, England) with addition of sterile pine acycles, at 25 °C with 12 h photoperiod of UV-light for a period of 49 d, for the production of reproductive structures (pycnidia). Once the pycnidia were formed, the color, shape, and presence (or not) of conidial septa were observed with an Olympus CX23 optical microscope (China), magnification of 40x(5)(11). In addition, the length and width of 30 conidia per isolate were recorded with the Dinno Capture 2.0 program.
2.3 Molecular identification
The colonies used were grown in PDA medium, incubated for five days at 25 °C in the dark, and the genomic DNA was extracted by Quick-DNA™ Fungal/Bacterial Mini-prep Kit (Zymo Research, USA), following the manufacturer's instructions. The DNA concentration of the samples was determined using a Nanodrop 2000 Thermo Scientific spectrophotometer, adjusting the concentration to 25 ng/μl. Amplification was performed by PCR with primers RPB2bot6F and RPB2bot7R from the partial region of the second major subunit of RNA polymerase II (RBP2)(11). Amplification conditions consisted of an initial denaturation at 94 °C for 4 m, followed by 10 cycles at 94 °C for 20 s, at 58 °C for 48 s, and at 72 °C for 45 s, and 25 cycles at 94 °C for 20 s, 56 °C for 40 s and 72 °C for 45 s; lastly, a final extension of 10 m at 72 °C. PCR amplification was carried out in a Peltier PTC-100 thermocycler (USA). PCR products (500 bp) were confirmed on a 1.5% agarose gel and sent for sequencing to Macrogen Inc., Korea. The obtained sequences were edited and manually corrected using the MEGA program version 10(12) and compared with deposited in GenBank through the BLAST search tool(13).
2.4 Pathogenicity tests
The “Queen Dream” hemp variety was used for the pathogenicity test, an American variety obtained by Blue Forest Farm with a 0.22% THC content(14).
The 30-day-old plants were transplanted into 0.8-liter pots with GrowMix® Multipro substrate. The three isolates grown in PDA for 5 d at 25 °C were used for inoculation (Table 1). Fifteen plants were inoculated per isolate, plus 12 plants for the non-inoculated control. The inoculation consisted of making a wound with a sterile toothpick, at the height of the cotyledons, and subsequent placement of a mycelium disk of 5 millimeters in diameter. Sterile PDA media discs were also placed in the control treatment plants. All discs were fixed to the wound with parafilm(5) and the plants were covered for 48 h with plastic bags previously moistened with sterile distilled water. The incidence of the disease was evaluated from 7 to 14 days post-inoculation under conditions of 25 ºC and a photoperiod of 16 h.
2.5 Reisolation
Once the symptoms and signs were observed, the pathogen was reisolated from the inoculated plants. Three symptomatic plants were taken per treatment for this purpose; pycnidia were extracted from the wounds with the help of a disinfected dissection needle and placed in Petri dishes with PDA medium and incubated in the dark at 25 ºC. The obtained colonies and conidia were characterized by morphology and compared with the characteristics of the initial isolates.
3. Results
3.1 Morphological identification
Three days after incubation, a whitish aerial mycelium with a cottony texture was observed, with very faint yellow pigment production from the center, observable on the back of the plate (Figures 1a and 1b). At eight days of incubation, the colonies presented a dense aerial mycelium with white-grayish colors observable from the front and olive green on the reverse of the plate (Figures 1c and 1d). At 16 days, the colonies presented a compact dark-colored mycelium, ranging from gray to black on the front and olive green to black on the back (Figure 1e and 1f).
After 49 days of incubation, it was possible to observe the presence of pycnidia with abundant hyaline conidia with smooth edges and without septa, in shapes ranging from ellipsoidal to spindle-shaped (Figure 2). The conidia sizes (n=30) of Cluc F, Migues 1 and Migues 14 averaged 16.6 μm (14.1-19.7) × 8.3 μm (7.1-9.7), 17.1 μm (14.4-19.8) × 8.4 μm (7.1-9.5), and 17.2 μm (14.4-19.7) × 8.4 μm (7.3-9.8), respectively.
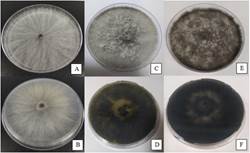 Figure 1
Isolate colony of N.
parvum. on the front and back of the plate in PDA medium after 3 (A-B), 8
(C-D) and 16 (E-F) days of incubation
Figure 1
Isolate colony of N.
parvum. on the front and back of the plate in PDA medium after 3 (A-B), 8
(C-D) and 16 (E-F) days of incubation
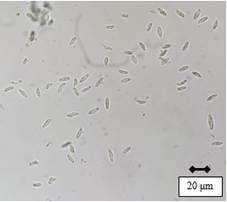 Figure 2
Conidia of N. parvum
observed under optical microscope with 40× magnification
Figure 2
Conidia of N. parvum
observed under optical microscope with 40× magnification
3.2 Molecular identification
The sequences Cluc F, Migues 1 and Migues 14 were deposited in the Gen-Bank (access number: OQ621792, OQ621793 and OQ621794, respectively). These showed 100% identity with N. parvum (MT592407.1) for the partial sequence of the second major subunit of RNA polymerase II.
3.3 Pathogenicity and reisolation test
Seven days after the inoculation, a brown darkening of the stem could be observed upwards from the inoculation area of the plant. As the disease progressed, it caused widespread wilting in the plant. At 14 days, black pycnidia developed on the stem wound (Figure 3). The incidence of disease was 86.6%, 73.3% and 60% in plants inoculated with Migues 14, Culc F and Migues 1, respectively. N. parvum was reisolated from symptomatic plants from stem pycnidia with characteristics consistent with the isolates used. Plants inoculated with PDA discs did not exhibit symptoms or signs of disease (Figure 3).
 Figure 3
Plants
inoculated with Migues 1 isolate of N. parvum showing generalized
necrosis (A), plant with pycnidia on stem wounds (B), healthy control plants (C-E)
Figure 3
Plants
inoculated with Migues 1 isolate of N. parvum showing generalized
necrosis (A), plant with pycnidia on stem wounds (B), healthy control plants (C-E)
4. Discussion
The study consisted of identifying and determining the pathogenicity of three isolates of Neofusicoccum obtained from plants with symptoms of stem canker. The isolates exhibited the phenotypic characteristics of the colonies and conidia described by Phillips and others(10) for Neofusicoccum spp.
During colony observation, it was possible to verify the production of yellow pigment, a recently observed characteristic in isolates of N. parvum according to a study carried out by Abdollahzadeh and others(15).
Values obtained of conidial size were within those presented by Phillips and others(10) for N. parvum 17.1 μm (12-24 μm) × 5.5 μm (4-10 μm), also similar to those of Delagado-Cerrone and others(7), 18.61 ± 2.33 × 7.38 ± 1.07. As for the characteristics of the conidia, they coincide with the description of Phillips and others(10), although no darkening of the conidia was observed over time, nor the presence of septa. These characteristics were not observed either in the isolates of Alberti and others(4) and Delagado-Cerrone and others(7), but may or may not occur within the species due to their high variability.
By analyzing the sequences of the partial region RBP2 N. parvum was identified. This region was also used to differentiate species from the complex N. parvum/N. ribis.(11).
The results of the pathogenicity tests showed that the isolates caused disease symptoms(2) in cannabis plants.
In Uruguay, N. parvum infects several plant species, but symptoms in cannabis had not been recorded. Thus, this study confirms N. parvum causing stem canker in cannabis, for the first time in Uruguay. Knowing the causal agent of the disease in hemp will allow the implementation of management strategies to reduce crop losses.
5. Conclusions
The three studied isolates were identified as N. parvum and pathogenicity in cannabis was proved. This is the first report of N. parvum causing stem canker disease on cannabis in Uruguay.
References
1. Sector cannabis en Uruguay [Internet]. Montevideo: Uruguay XXI; 2021 [cited 2023 Jun 12]. 28p. Available from: https://bit.ly/3oZOlGZ
2. Rousserie G, Collazo D. Prospección de plagas, hongos y oomicetos asociados a enfermedades en Cannabis sativa L. en el sur de Uruguay [grade’s thesis]. Montevideo (UY): Universidad de la República, Facultad de Agronomía; 2022. 174p.
3. Phillips AJL, Alves A, Abdollahzadeh J, Slippers B, Wingfield MJ, Groenewald JZ, Crous PW. The Botryosphaeriaceae: genera and species known from culture. Stud Mycol. 2013;(1):51-167. Doi: 10.3114/sim0021.
4. Alberti I, Prodi A, Nipoti P, Grassi G. First report of Nepfusicoccum parvum causing stem and branch canker of Cannabis sativa in Italy. J Plant Dis Prot. 2018;125(3):511-3. Doi: 10.10d07/s41348-018-0174-4.
5. Feng C, Villarroel-Zeballos MI, Ficheux PF, Zima H, Dhillon BDS, Correll JC. First report of Neofusicoccum parvum causing dieback and canker disease on hemp in the United States. Plant Dis. 2020;104(11):2754-3089. Doi: 10.1094/PDIS-03-20-0486-PDN.
6. Sessa L, Abreo E, Bettucci L, Lupo S. Botryosphaeriaceae species associated with wood diseases of stone and pome fruits trees: symptoms and virulence across different hosts in Uruguay. Eur J Plant Pathol. 2016;146(3):519-30. Doi: 10.1007/s1065 8-016-0936-4.
7. Abreo E, Martínez S, Bettucci L, Lupo S. Characterization of Botryosphaeriaceae species associated with grapevines in Uruguay. Australas Plant Pathol. 2013;42(3):241-9. Doi: 10.1007/s1331 3-013-0200-8.
8. Delgado-Cerrone L, Mondino-Hintz P, Alaniz-Ferro S. Botryosphariaceae species associated with stem canker, die-back and fruit rot on apple in Uruguay. Eur J Plant Pathol. 2017;146(3):637-55.
9. Pennycook S, Samuels G. Botryosphaeria and Fusicoccumspecies asociated whit ripe fruit rot of Actinidia deliciosa (kiwifruit) in New Zealand. Mycotaxon. 1985;24(24):445-58.
10. Phillips A, Alvez A, Abdollahzadeh J, Slippers B, Wingfield J, Groenewald J, Crous P. The Botryosphaeriaceae: genera and species know from culture. Stud Mycol. 2013;76:51-167. Doi: 10.3114/sim0021.
11. Pavlic D, Slippers B, Coutinho TA, Wingfield MJ. Multiple gene genealogies and phenotypic data reveal cryptic species of the Botryosphaeriaceae: a case of study on the Neofusicoccum parvum.N. ribis complex. Mol Phylogenet Evol. 2009;51(5):259-68. Doi: 10.1016/j.ympev.2008.12.017.
12. Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol. 2018;35(6):1547-49. Doi: 10.1093/molbev/msy096.
13. Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW. GenBank. Nucleic Acids Res. 2013;41(D1):D36-42. Doi: 10.1093/nar/gks1195.
14. Gobi Labs. Analytical Report [Internet]. Wheat Ridge: Gobi Labs; 2020 [cited 2023 Jun 12]. 1p. Report no.: P-3974857-V1. Available from: https://bit.ly/45W3AS3
15. Abdollahzadeh J, Zare R, Phillips A. Phylogeny and taxonomy of Botryosphaeria and Neofusicoccum species in Iran, with description of Botryosphaeria scharifii. Nov. Mycologia. 2013;105:210-20. Doi: 10.3852/12-107.
Author notes
elvisjoa14@gmail.com
Additional information
Transparency
of data: Available data: The entire data set that supports the
results of this study was published in the article itself.
Author
contribution statement: SE:
morphological identification, pathogenicity test, interpretation of results,
and elaboration of the manuscript; GRP: co-author of the study and manuscript
revision; CD: isolation of the fungus and molecular identification; RG:
isolation of the fungus and molecular identification; SPE: study tutor,
interpretation of results, revision, and edition of the manuscript.
Editor: The following editor approved this article:
Gustavo Giménez (https://orcid.org/0000-0001-7176-185X) Instituto Nacional de
Investigación Agropecuaria (INIA), Canelones, Uruguay
Alternative link
https://agrocienciauruguay.uy/index.php/agrociencia/article/view/1172/1458 (pdf)









