1.
Introduction
The Leguminosae (Fabaceae) is the third-largest plant family among angiosperms that comprises 20,000 species and 750 genera(1)(2)(3)(4). Legumes establish a symbiotic association with Gram-negative soil bacteria known as rhizobia(1)(5) that results in the development of nodules in roots and stems, where atmospheric nitrogen is reduced (fixed) to ammonia, that is assimilated by plants into organic compounds. The family of legumes is divided into three subfamilies that include mostly plants adapted to tropical regions, Cesalpinoidea, Papilinoidea and Mimosoidea; within the former one, only 11 genera form nitrogen-fixing nodules(1)(6). Rhizobia are clustered within the Alpha and Betaproteobacteria(7), intermingled with non-symbiotic, photosynthetic and plant pathogenic genera and distributed in at least 14 genera(7). Rhizobia vary in specificity as well as legumes, while some of them nodulate several hosts, others nodulate only one or two species. Furthermore, several hosts are nodulated by several rhizobia, including both Alpha and Betaproteobacteria. In other cases, the rhizobial strain-host genotype interaction is highly specific, to the point that two closely related species of plants are nodulated by different species of rhizobia(7).
Legume nodules induced by rhizobia also might be co-inhabited by a set of various Gram-positive and Gram-negative bacteria, some of them unable to induce nodule development(1)(6)(7)(8). These endophytic bacteria can promote plant growth through many different mechanisms and they can do so in symbiosis or as free-living cells. Plant Growth-Promoting Bacteria (PGPB) may act directly by increasing the availability of nutrients such as P solubilization or N2 fixation, by influencing plant hormone levels, or indirectly by attenuating the effects of pathogens(9)(10), increasing in this way host fitness. In line with this, rhizobia also may act as PGPB in non-legume cash-crops such as rice or wheat, which turn out to be some of the best beneficial interaction between endophytic rhizobia and plants(11).
Both tree legumes and actinorricic plants contribute to increasing N in soils, which is the main reason to consider such systems as a sustainable strategy to improve the productive potential of many different agroecosystems(12), to manage silvicultural systems or to prevent soil degradation and N depletion as well. Many legume trees contribute N to tropical wetlands and rainforests, and because of this, it is necessary to have a better understanding of such systems to improve the use and conservation of natural resources(13). Additionally, such systems might be exploited in reforestation as well as in land restoration(14). A good example of such systems is the Acacia sensu lato species, highly tolerant to drought, due to their large root systems that not only lead plants to explore more soil but also to access deeper layers of water. Vachellia sp. are fast-growing plant species pioneers in extreme environments due to their nitrogen-fixing ability, and also to other symbioses they establish with fungi (e.g. endo and/or ectomycorrhiza), which are critical for their role in poor and eroded soils(14). Many legumes might be relevant tools to increase soil organic matter in such systems. Species of Poecilanthe grow in northeast Brazil, in arid areas where water is the main limiting factor. In such ecosystems, a subsistence agriculture mainly based in extensive livestock prevail, where pastures of native legumes are among the most important sources of cow food.
Considering the potential of tree legumes, several studies have been conducted in South America(12)(13)(14)(15)(16)(17)(18)(19)(20)(21)(22)(23)(24)(25)(26)(27)(28)(29)(30). They consisted in the isolation of rhizobia and the study of nodule morphology, biochemical and genetic characteristics, systematics and phylogenetic relationships of bacterial symbionts, as well as their ability to nodulate and fix N. However, there are not very many reports regarding species like P. parviflora (Lapachillo), an endemic tree to South America, Vachellia caven (Espinillo) and E. contorsiliquum (Timbó). Regarding the genus Poecilanthe, there are only a few reports of bacteria isolated from nodulated plants growing in Brazil(13)(30). Thus, it is of great interest to study the rhizobia as well as other bacteria (nodule microbiome) associated with Poecilanthe sp., Vachellia sp., and Enterolobium sp. in South-American ecosystems, considering their role in nature, particularly regarding legumes growth and development(31).
The aim of this work was to isolate and characterize, based on microbiological as well as molecular markers, bacteria within nodules of three native trees species from South America: Poecilanthe parviflora Benth, Vachellia caven (Molina) Seigler & Ebinger, and Enterolobium contortisiliquum (Vell.) Morong. Such information might lead to make a better use of the potential of these native legumes in complex environments.
2. Materials and methods
2.1 Source of bacterial endophytes and biological
material
Composite samples of soil were collected from places located under the canopy of P. parviflora plants. Nodules were collected from roots of plants growing within the marginal rain forest of Martín García Island, a protected area located at 34°11′19″S, 58°15′00″W; 14 m.a.s.l (meters above sea level), Buenos Aires, Argentina, belonging to the Delta and Islas del Paraná ecoregion(32). Nodules were maintained in hermetic containers until they were processed, which occurred within 48 h. Collected plants were transferred and maintained in 3 L pots filled with soil from Martín García Island and vermiculite at a 1:1 ratio. Pots were watered every other day with distilled water.
Nodules collected from plant roots were surface sterilized by immersing them 2 min. in ethanol 50% followed by 2 min. in sodium hypochlorite 4 g L-1. Then, they were rinsed at least 10 times with sterile distilled water. Sterilization of nodule surfaces was checked by rolling them on plates filled with: a) Yeast Extract Mannitol Agar (YMA)(33) K2HPO4 0,5 g L-1, MgSO4. 7 H2O 0,2 g L-1, NaCl 0,1 g L-1, CO3Ca 1 g L-1, yeast extract 1 g L-1, mannitol 10 g L-1, congo red (1/400) 10 ml, Agar 15 g L-1, Ph 6.8; b) Cetrimide agar(34) (gelatin peptone 20 g L-1, magnesium chloride 1.4 g L-1, potassium sulfate 10 g L-1, cetyltrimethylammonium bromide 0.3 g L-1, Agar 15 g L-1), and c) Nutrient agar(35) (peptone 5 g L-1, meat extract 3 g L-1, sodium chloride 8 g L-1, Agar 15 g L-1).
Isolations were made according to Vincent(33) using the described YMA and Nutrient agar media and inoculated plates were incubated at 28 °C. After 7-day-incubation period colonies were subcultured until pure cultures of the isolates were obtained. Then, isolated bacteria were grown in YEM broth(33), until saturation and aliquots were mixed with glycerol to make a final concentration of 10% in stocks that were maintained at -80 °C.
V. caven and E. contorsiliquum nodule bacteria were obtained as described before, from nodules developed by inoculated plants. Seeds of these species were surface scarified and sterilized with sulfuric acid for 15 min., rinsed with sterile water and germinated in 10% water-agar in Petri dishes in the dark. Seedlings were transplanted into pots with sterile vermiculite and inoculated with 1 ml of soils decimal dilution in sterile water. Soils were collected at 24°48’52.3’’S 65°29’59.6’’W, 1.300 m.a.s.l. for V. caven. and at 24°35’18.7’’S 65°02’50’’W, 750 m.a.s.l. for E.contorsiliquum, from the dry Chaco ecoregion(32), Salta, Argentina. Stock cultures of isolated bacteria were made and kept as described before. Also, reference strains B. japonicum E109, SEMIA 5079, B. elkanii SEMIA 5019 and SEMIA 587, as well as the type strain Mesorhizobium Chacoense Pr5 (=LMG19008=CECT5336) isolated from Prosopis alba(36) at the dry Chaco ecoregion(32), Chancaní Reserva, Córdoba, Argentina, were included in this work.
2.2 Microbiological characterization
All isolates were observed with the light field optical microscope (1000X) after Gram staining(37). Alkalinization or acidification of the culture medium was determined in non-buffered YEM broth supplemented with bromothymol blue, after 10 days of growth, at 28 °C in a rotary shaker at 250 rev. min-1. Isolates were classified as developing a strong, moderately or slightly acid reaction based on the yellow colour developed, while neutral and basic reactions were associated with green and blue colours, respectively.
Bacterial isolates were grown for 10 days at 28 °C on NBRI-P (National Botanical Research Institute's phosphate)(38) medium (glucose 10g L-1, Ca3PO4 5 g L-1, MgCl2.6H2O 5 g L-1, KCl 0,2 g L-1, (NH4)2SO4 0.1g L-1, SO4Mg.7 H2O, Agar 20g L-1) to evaluate their ability to solubilize inorganic phosphorous that was determined by the development of a transparent halo around bacterial colonies.
The antimicrobial activity of isolates was assayed by co-culturing them with two model plant pathogens, Stemphylium lycopersici CIDEFI 216, the causal agent of grey leaf spot(39), and Fusarium graminearum SP1, the etiological agent of head blight(40). These two pathogens have a wide host range, representatives of them have been reported to provoke diseases on legumes and are used as model systems to study plant-microbe interactions in our laboratory. Both fungi were grown on PDA (Potato Dextrose Agar, potato 250 g L-1, dextrose 20 g L-1, agar 20 g L-1)(41) medium, for 3 (F. graminearum) or 10 (S. lycopersici) days at 25 °C in the darkness. Antimicrobial activity was evaluated by placing bacteria on PDA medium in two equidistant parallel streaks. After an incubation period of approximately 7 days at 25 °C, a disc of the fungus was placed at the center of each Petri dish. Control treatment consisted of plates inoculated with fungi in the absence of bacteria. All plates were sealed with parafilm and incubated at 25 °C. The experiment finished within 3 (F. graminearum) or 10 (S. lycopersici) days.
The inhibitory activity of bacteria was estimated by measuring mycelial growth on the surface of the Petri dishes using the software Image tool 3.0 (Image Tool Software Copyright)(42). The percentage of inhibition (InC) was calculated with the formula: % InC = [(control mycelial growth area - treatment mycelial growth area)/control mycelial growth area]*100(43). Bacterial isolates ability to solubilize inorganic phosphorous and their antagonistic effect against S. lycopersici and F. graminearum were evaluated in three independent experiments. The collected data were subjected to a one-way analysis of variance (ANOVA), means were compared by the Tukey test. All statistical analyses were performed using Infostat version 2015(44).
2.3 Molecular analysis
2.3.1 DNA
extraction and genetic diversity analysis
Isolates DNA was extracted using the Wizard® Genomic DNA Purification Kit (Promega)(45). Briefly, isolated bacteria were cultured in liquid media until a cell concentration of 1×109cells mL−1 was obtained. Aliquots of 5 mL were used to extract DNA, whose quality and quantity was checked by electrophoresis in 0.7% agarose gels stained with ethidium bromide that included a molecular marker(46).
Endophytic bacterial isolates were fingerprinted using a BOX-PCR with the universal BOXA1R primer (5′-CTACGGCAAGGCGACGCTGACG-3′). PCR contained 1x amplification buffer (Inbio Highway), 2.5mM MgCl2 (Inbio Highway), 50 pmol BOXA1R primer, 2 mM each dNTP (Inbio Highway), 50-100 ng of genomic DNA, and 0.6 U Taq DNA polymerase (Inbio Highway) in a 15 ul volume. Reactions were performed in a PTC-1152 Mini Cycler (MJ Research) programmed as follows: an initial step at 94 °C for 7 min, followed by 35 cycles at 94 °C for 1 min, 53 °C for 1 min, and a final step at 65 °C for 8 min, and a final cycle at 65 °C for 16 min(47). The PCR amplicons were separated on 1.5% agarose gels, stained with ethidium bromide.
The sizes of the fragments were normalized according to the MW of the DNA markers (λ HindIII marker, Invitrogen). Fingerprints were analyzed in a binary data matrix, scoring 0 for the absence and 1 for the presence of bands. A multivariate analysis was carried out using Past3 software(48).The Dice similarity index was used to create a similarity matrix that was used to generate a dendrogram using the Unweighted Pair Group Method with Arithmetic Mean (UPGMA) algorithm(39). All those bacterial cultures that had a unique fingerprint were selected for 16S rDNA analysis.
2.3.2 16S rDNA
amplification and sequencing
Bacterial isolates were identified using a partial sequence of the 1500 bp 16SrDNA. Such fragments were amplified by PCR in a thermocycler (MinicyclerTM, MJ Research Inc., Waltham, MA, USA), using primers 27F (5´-AGAGTTTGATCMTGGCTCAG-3´) and 1492R (5´-ACGGTTACCTTGTTACGACTT-3´) as described by Reysenbach and others(49). PCR products were purified as described by Sambrook and others(50), precipitated and sequenced at MACROGEN Inc. (Seoul, South Korea). Sequences were analysed and trimmed with Geneious R9 software version R9.0.5, Biomatters(51). Sequences were analysed by Basic Local Alignment Search Tool (BLAST) against 16S rDNA sequences of type strains.
The phylogenetic analysis was conducted using Geneious R9 software(51). Sequences were aligned using the default parameters of the ClustalW algorithm (gap opening penalty 15, gap extension penalty 6.66)(52). Phylogenetic analysis was performed using the genetic distance model as described by Tamura- Nei using the neighbor-joining method(53).
3. Results and discussion
3.1 Isolation of endophytic bacteria from nodules of
tree legumes from Argentina
The isolates collected from nodules of native legume trees included in this work are presented in Table 1. All P. parviflora isolates were obtained from root nodules belonging to marginal rain forest plants, which was the only environment in the Martín García Island, among the three sampled sites (riparian forest, marginal rain forest and the coronillo forest). We found P. parviflora plants with functional indeterminate nodules whose interior were red, suggesting they contain leghemoglobin and are fixing nitrogen. In the riparian and the coronillo forests, samples of P. parviflora plants had no nodules or non-functional ones. This may be due to the different soil texture classes, frequent flooding of these soils and/or variations in legume species composing the flora at the different vegetative units, associated to the environmental gradient of the island(54)(55), which might influence and/or determine the pools of resident bacteria in the soil.
Among isolated bacteria, 15 were obtained from P. parviflora, 6 from V. caven and 1 from E. contortisiliquum. Interestingly, the isolates from P. parviflora were recovered, some of them on one media and the other ones on other media, suggesting that nodules probably harbour a diverse array of bacteria.
3.2 Microbiological characteristics of the isolates
Four
bacteria among those collected from P. parviflora and the E. contorsiliquum isolate alkalinize the
culture medium, while 1 from M. chacoense, 7 from P. parviflora and all 6 isolated from nodules of V. caven acidified the media. However,
while some of them generated a slight acid reaction, others generated a strong
one on the same media. The other 4 bacteria isolated from P. parviflora did not alter the pH of the medium.
The main microbiological characteristics of the isolates are presented in
supplementary material (Table S1). It is known that while Bradyrhizobium sp.
alkalinize the YEM broth, Ensifer
(Sinhorhizobium) acidified it and Mesorhizobium
species activity on the culture broth is dependent on the carbon source of
the media(36).
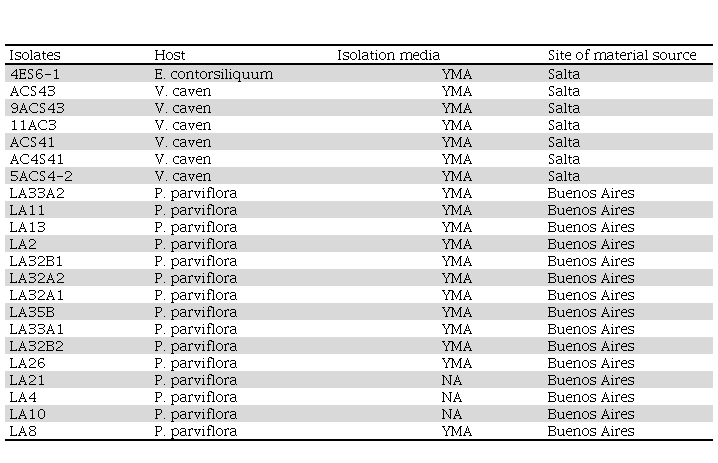
Table 1
Bacterial isolates obtained from nodulated
plants and control strains used in our experiments. Host species and the media
used to isolate bacteria as well as the site of material source (soil sample or
nodule plant collected in situ) used for the bacterial isolation are presented.
YMA: Yeast Extract Mannitol Agar supplemented with congo red; NA: Nutrient
Agar. The isolates were named according to the host of origin E: Enterolobium, AC: Acacia (Vachellia) caven
and LA: Lapachillo. Numbers before and after the
letters indicate the number of the isolate, except for the denominations as an
"S" followed by number (e.g. S43), which
indicate different soil sites.
|
Isolates
|
Host
|
Isolation media
|
Site of material source
|
|
4ES6-1
|
E. contorsiliquum
|
YMA
|
Salta
|
|
ACS43
|
V. caven
|
YMA
|
Salta
|
|
9ACS43
|
V. caven
|
YMA
|
Salta
|
|
11AC3
|
V. caven
|
YMA
|
Salta
|
|
ACS41
|
V. caven
|
YMA
|
Salta
|
|
AC4S41
|
V. caven
|
YMA
|
Salta
|
|
5ACS4-2
|
V. caven
|
YMA
|
Salta
|
|
LA33A2
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA11
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA13
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA2
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA32B1
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA32A2
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA32A1
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA35B
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA33A1
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA32B2
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA26
|
P. parviflora
|
YMA
|
Buenos Aires
|
|
LA21
|
P. parviflora
|
NA
|
Buenos Aires
|
|
LA4
|
P. parviflora
|
NA
|
Buenos Aires
|
|
LA10
|
P. parviflora
|
NA
|
Buenos Aires
|
|
LA8
|
P. parviflora
|
YMA
|
Buenos Aires
|
| | | | |
We first screened the isolates to identify their capacity to inhibit S. lycopersici and F. graminearum growth (data no show). B. japonicum SEMIA5079 and isolates 4ES6-1, 5ACS4-2, LA33A2 and LA8 inhibited fungal growth at values <10%. Based on these results we run additional experiments with selected isolates and reference strains that inhibited mycelial growth ≥10% of any of the pathogenic fungi assayed. Among isolates of P. parviflora, V. caven and E. contorsiliquum analyzed (Figure 1), 11 of them inhibited the growth of F. graminearum and/or S. lycopersici within a range of 19% to 72% and 41% to 95%, respectively. Both fungal pathogens F graminearun and S. lycopersici were inhibited by the following 5 isolates of P. parviflora: LA4, LA10, LA 11, LA21 and LA26. The former pathogen was inhibited in a 34%, 23%, 68%, 72% and 26%, and the latter one in an 82%, 80%, 91%, 95% and 43%, respectively. Also, isolate 9ACS43 obtained from V.caven inhibited F. graminearum and S. lycopersici in a 19% and 41%, respectively. The commercial strain B.elkanii 5019 behaved like the isolates described, inhibiting F. graminearum and S. lycopersici by a 23 and 49%, respectively (Figure 1).
Two isolates collected from V. caven (ACS43 and 11AC3) and three from P.parviflora (LA13, LA32B2, LA35B) as well as M. chacoense only inhibited the growth of S. lycopersici or F. graminearum (Figure1) with values on S. lycopersici of 44% (LA13) and 64% (M. chacoense), and 28% (ACS43), 29% (LA32B2), 36% (LA35B) and 52% (11AC3) for F. graminearum. Among the isolates that proved to have a considerable capacity to inhibit fungal growth, only isolate 11AC3 that was collected from nodules of V.caven was able to solubilize phosphorus, suggesting that it might be a promising plant growth-promoting bacteria that not only control plant pathogens but also might help plants to access more P, that most of the time is in unavailable forms in the soil.
3.3 Genetic
studies: diversity and identification of endophytic bacteria
Molecular biology provided researchers with powerful methods for the typing and genetic identification of organisms. Such information is crucial and is used to perform phylogenetic analysis, since universal conserved gene sequences such as the 16S rDNA are used. DNA-based methods have become increasingly important to identify and characterize rhizobia, in particular, phylogenetic analyses of sequences of the 16S ribosomal rDNA (rRNA) gene, a range of “housekeeping” genes and genes involved in symbiosis have been used and proposed as a "standard approach" for the final identification of organisms. Concomitantly, the 16S rDNA gene sequence on its own turn out to be a reliable tool mainly for the preliminary identification of microorganisms(56). However, several gene sequences like those of the nif and nod genes involved in N2 fixation and legume host specificity are often carried on plasmids or symbiotic islands in many nitrogen-fixing bacteria, and therefore, might be horizontally transferred between different species within a genus and less frequently across genera(57)(58); because of this their relevance or role in identification has to be cautiously considered.
Based on the reasons described we compared the genome fingerprints generated using the BOXAR1 primer of isolated bacteria with those of B.Diazoeficiens (SEMIA 5080), B. japonicum (E109 and SEMIA 5079), B. elkanii (SEMIA 5019 and SEMIA 587), and M. Chacoense type strain using the BOXAR1 primer PCR (Figure 2).
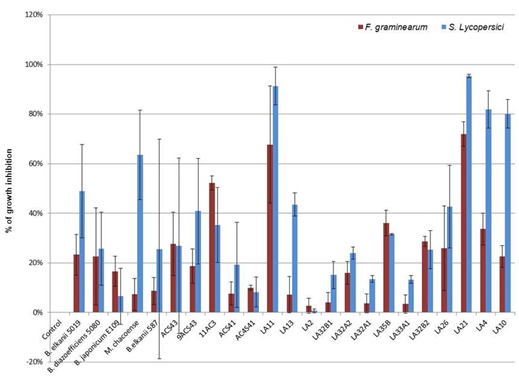 Figure 1
Growth inhibition
in vitro of fungal pathogens F. graminearum and S.
lycopersici by 18 bacterial isolates. B. elkanii 5019
and 587, B.diazoefficiens SEMIA5080, B. japonicum E109 and M. chacoense
were included as controls
Figure 1
Growth inhibition
in vitro of fungal pathogens F. graminearum and S.
lycopersici by 18 bacterial isolates. B. elkanii 5019
and 587, B.diazoefficiens SEMIA5080, B. japonicum E109 and M. chacoense
were included as controls
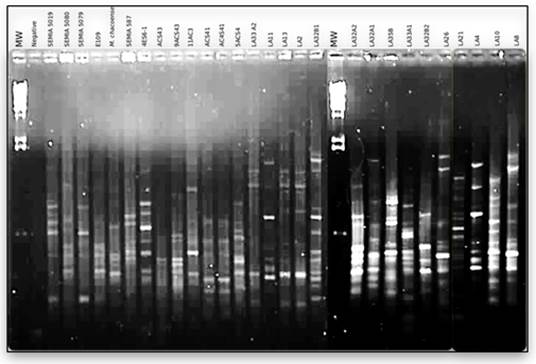 Figure 2
BOXPCR
fingerprints of bacteria isolated from nodules of from P. parviflora, V. caven and E. contorsiliquum
Figure 2
BOXPCR
fingerprints of bacteria isolated from nodules of from P. parviflora, V. caven and E. contorsiliquum
Isolates and type strains were grouped into 4 major clusters (Figure 3) at a ~0.35 similarity level, two of them only containing isolates of P. parviflora. Cluster I was formed by isolates LA11, LA4 and LA21, and Cluster IV included isolates LA33A2, LA13 and LA2. Clearly, these two groups of bacteria were quite different. The other 16 isolates were grouped within cluster II and III. Cluster II included isolates of P. parviflora LA33A1, LA10 and LA32B1 that were distributed into 3 different subgroups, LA26 and LA8 were together and in a different subgroup than isolate 5ACS4-2 of V.caven, which also was different from V. caven 9ACS43 and ACS43, that were grouped at a ~0.7 similarity level. Cluster III included isolates of V. caven, P. parviflora, E. contorsiliqum, M. Chacoense, B. elkanii, B. japonicum and B. diazoeficiens, so most probably, as shown below, these bacteria were representatives of nitrogen-fixing ones. At a 0.6 similarity level, the isolate of P. parviflora LA32A1 was grouped with bacteria isolated from E. contorsoliquum in a different subgroup, apart from isolates LA35B, LA32A2 and LA32B. The isolate 11AC3 of V. caven differed from ACS41 or AC4S41, although additional genetic studies should be performed to accurately establish similarity between the isolates. Cluster IV included two isolates that had an identical fingerprint and therefore might be siblings of the same strain, while the other isolate proved to be quite diverse. In any case, organisms within this cluster are the more diverse compared to the other bacteria. It is evident that even though isolates were collected from the same hosts, they were genetically quite diverse, and even nitrogen fixing bacteria might belong to different rhizobial species as well.
Based on the BOX dendrogram we proceeded to identify the most diverse bacteria by amplifying and sequencing their 16S rDNA gene. The BLAST analysis of these sequences suggested that root nodule endophytic bacteria belonged to 10 genera that were either within the Alphaproteobacteria, Gammaproteobacteria, Firmicutes or Actinobacteria (Figure 4).
Within nodules we isolated bacteria that based on the 16S rDNA sequence were clustered in 4 main groups (Figure 4). Since they were isolated from nodules, they might either be nitrogen-fixing bacteria, plant growth-promoting bacteria that live within nodules, or opportunistic saprophytic bacteria that alone or by interacting with the plant or other microbes, provide plants with a biological advantage for the ecological niche. One group of bacteria consisted in Gammaproteobacteria and included isolates LA8 and LA10 that were isolated from nodules of P. parviflora. These bacteria belonged to the families Rhodanobacteraceae (Dyella sp.) and Pseudomonadaceae (Pseudomonas sp.), with 99.86% and 100% of identity, respectively. This same cluster also included isolate 11AC3, that was collected from nodules of A. caven and belonged to Enterobacteraceae, being an Enterobacter sp. However, the sequence identity was too low, therefore, more sequences or a larger 16SrDNA sequence should be analyzed to accurately identify the isolate.
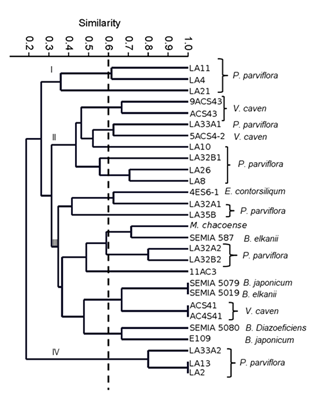 Figure 3
Dendrogram
generated by UPGMA cluster analysis using the Dice similarity coefficient based
on the BOXAR1 fingerprint of studied isolates
Figure 3
Dendrogram
generated by UPGMA cluster analysis using the Dice similarity coefficient based
on the BOXAR1 fingerprint of studied isolates
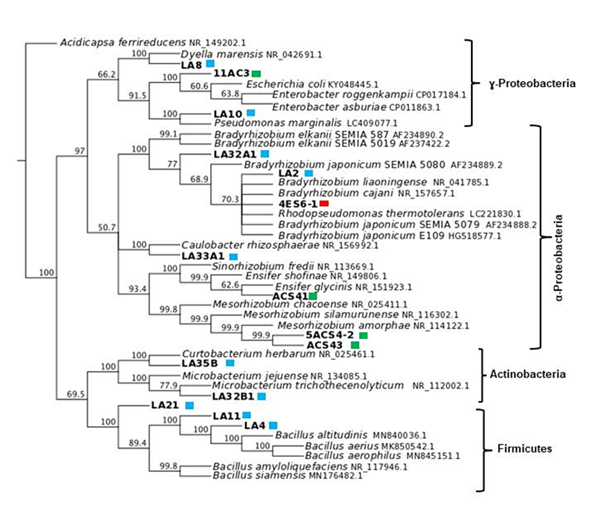 Figure 4
Phylogenetic
relationship of the isolates based on aligned sequences of the 16S rDNA gene.
Sequences accession number representing the genera is on the tree. Isolates
from the same legume species are indicated as coloured
squares: blue for P. parviflora,
green for V. caven,
and red for E. contorsiliquum.
Acidicapsa ferrireducens
was the outgroup
Figure 4
Phylogenetic
relationship of the isolates based on aligned sequences of the 16S rDNA gene.
Sequences accession number representing the genera is on the tree. Isolates
from the same legume species are indicated as coloured
squares: blue for P. parviflora,
green for V. caven,
and red for E. contorsiliquum.
Acidicapsa ferrireducens
was the outgroup
Among isolated bacteria, the blast analysis showed that at least 6 were nitrogen-fixing bacteria that belonged to Alphaproteobacteria. These were isolates LA2 from P. parviflora; 4-ES6-1 from E. contorsiliquum; isolates ACS41, 5ACS4-2, ACS43 that were collected from nodules of A. caven; and isolate LA32A1, whose 16SrDNA sequence, unfortunately, had a too low similarity level, indicating that we need to go back to more precisely identify the isolate. These bacteria, except for the latter one, were clustered separately with a high bootstrap value of 100% and 93.4%. Furthermore, they are most probably representatives of Bradyrhizobium, Mesorhizobium and Ensifer since their sequences shared 99.66 % and 100% identity with type strains of these genera. It has already been reported that nodules induced by rhizobia might contain other bacterial taxa such as representatives of the Gammaproteobacteria. Pantoea agglomerans, Enterobacter kobei, Enterobacter cloacae, Leclercia adecarboxylata, Escherichia vulneris, Pseudomonas sp., Bacillus sp., Pseudomonas sp., Curtobacterium herbarum and Microbacterium sp., among others, which were frequent bacteria within nodules of Hedysarum mediterranean species(59)(60). This might be related to the nutrient level and/or particular environment of nodules that evidently harbour a wide variety of non-symbiotic bacteria, which could be either necessary partners or opportunistic competitors.
Other isolates were clustered within the Actinobacteria group, like LA32B1 and LA35B organisms, that were isolated from P. parviflora and identified one as Microbacterium and the other as Curtobacterium. Representatives of the Firmicutes LA4, LA11 and LA21 were identified as Bacillus species, since their 16SrDNA sequence was 100% identical to that of Bacillus species. Bacteria recovered from native legume trees live endophytically within nitrogen-fixing nodules induced by rhizobia, but might be either cooperative microorganisms, plant growth promoting bacteria or nitrogen fixing ones. It has already been described that Pseudomonas (Gammaproteobacteria) developed N2 fixing nodules on Robinia pseudoacacia(61) and Acacia confusa(62). Geobacillus (Firmicutes), Paenibacillus (Firmicutes) and Rhodococcus (Actinobacteria) also were described as rhizobial symbionts of Lotus corniculatus(63). These bacterial species had similar nodA gene sequences to Mesorhizobium representatives isolated from the same plants; most probably these bacterial species received these genes by lateral gene transfer from Mesorhizobium. Although such process might not be so frequent, the analysis should be done to clarify this issue. As reported by Saïdi and others(64), Pseudomonas, Enterobacter, Bacillus, Staphylococcus, Serratia, Stenotrophomonas and Xanthomonas have already been found within Vicia faba root nodules, though these bacteria failed to nodulate it and did not have nifH or nodC genes. However, they also cited reports of Alphaproteobacteria, Betaproteobacteria and Gammaproteobacteria that were isolated from root nodules as nodulating bacteria or nodule associated bacteria in a wide array of legumes. All this together suggests that legume nodulation may not be exclusively triggered by Alphaproteobacteria(8). Also, the other organisms living inside nodules provide plants with additional advantages that might be related to nutrition or plant health. This is not surprising considering that rhizobia free legumes like alfalfa can develop nodules even in axenic conditions(65), suggesting that legumes have preprogram sets of events that allow them to develop nodules. Probably, there are many more organisms than rhizobia alone that can trigger nodule formation in legumes(66).
Rhizobia isolates were clustered in three groups belonging to the families Bradyrhizobiaceae, Rhizobiaceae and Phyllobacteriaceae (Figure 4), among them, Mesorhizobium is closer to Sinorhizobium than to Bradyrhizobium, which is in agreement with previous reports(11). Isolates LA2, LA32A1 and 4ES61 that were isolated from P. parviflora and E. contorsiliquum, respectively, belonged to a cluster that included B. japonicum strains, suggesting that they most probably are Bradyrhizobium sp. Interestingly, B.Elkanii SEMIA 6403 (=BR 8205) has been recommended in Brazil to inoculate P. parviflora(67). Nodules of E. contorsiliquum contained a Bradhyrhizobium sp. that was characterized by an extremely slow growth rate and very small cells, such that they were hardly visible in the light microscope. This and the 16S rDNA sequence homology with the 16SrDNA sequence of B. liaoningense type strain (data not shown) suggested that it is most probably a B. liaoningense representative. In line with this, cultures of such isolate alkalinize the culture media. Isolates from P. parviflora belonged to Bradyrhizobium genus and, even though they were different, additional sequences should be examined to accurately establish their identity, particularly because there are other native legumes within the area suggesting that they might be promiscuous bacteria.
V. Caven was nodulated by Mesorhizobium strains since isolates 5ACS4-2 and ACS43 clustered with Mesorhizobium representatives with a high bootstrap value of 99.88 %. The 16S rDNA sequence of the isolated strains were 99.79 % and 100 % identical to that of Mesorhizobium strains. Unlike other genera of rhizobia, species within Mesorhizobium show relatively low sequence divergence at core loci, this is such that all Mesorhizobium type strains presented a high level of sequence homology(11)(68). Laranjo and others(11) grouped 30 Mesorhizobium species in four clusters, which as in this work showed differences between them. Furthermore, MLSA phylogeny analysis divided isolates in three main groups that contained M. plurifarium and M. chacoense, suggesting, in a way, the low resolution of the 16SrDNA gene sequences(11). They stated that lateral gene transfer, as well as duplication, might bring additional problems for phylogenetic studies within the genus Mesorhizobium. Because of this, the phylogenetic analysis of Mesorhizobium might disagree with broader phylogenies based on multiple core genes(68)(69). However, Laranjo and others(11) found that there is no gene transfer of core loci between some specific groups of Mesorhizobium.
Vachellia is a genus that includes promiscuous species that might be nodulated by several species of rhizobia(70). Our results confirmed this since isolate ACS41 was clustered with Sinorhizobim sp. (Ensifer sp.) at 100% bootstrap value and a 99.85% of sequence identity, suggesting that it belongs to the genus Ensifer. Interestingly, Ensifer representatives have been found to have the widest host range so far studied(71), and Vachellia species were nodulated by diverse and promiscuous Ensifer species(72)(73).
The rhizobia isolated for the native trees included in our study are in line with previous findings that stated that 59 Mesorhizobium species nodulate a wide array of different legumes(74): M. amorphae from Amorpha fruticosa(75), M. silamurunense from Astragalus membranaceus(76), and Mesorhizobium acaciaefrom Acacia melanoxylon R. Br.(77), among others. Beukes and others(70) studied Vachellia karroo root nodules that contained the genera Bradyrhizobium, Ensifer (32 isolates), Mesorhizobium (27 isolates), and Rhizobium, as well as Betaproteobacteria; and more recently, Pereira-Gómez and others(22) reported 8 V. caven isolates, affiliated to the genus Mesorhizobium. Thus, this diversity may contribute to the ecological success of Vachellia sp. as pioneer species, as previously noted, for example, for V. Karoo(70).
Both genera Ensifer and Mesorhizobium include several species that nodulate Acacia sensu lato species; this is in agreement with previous data reported for other African Acacias (i.e., Vachellia sp. and Senegalia sp.), which also are predominantly nodulated by isolates of Mesorhizobium, Rhizobium and Ensifer(70). Also, Cordero and others(14) found that V. macracantha, a legume tree distributed across neotropical regions, spreading from the Southern United States to Northern Argentina, was nodulated by Ensifer numidicus, Ensifer kummerowiae, Ensifer fredii, Ensifer americanus and Ensifer saheli. A range of genera in the Proteobacteria (most commonly Bradyrhizobium, Ensifer (Sinorhizobium), Mesorhizobium and Rhizobium) can form functional (N2 fixing) nodules on specific legumes, and for the Mimoseae tribe genera Acacia, Senegalia, Prosopis and Vachellia, and a complete list of legume —rhizobia symbiosis— is available in Andrews and Andrews(56).
4. Conclusions
South American native tree legumes V. caven and P. parviflora harbour a great diversity of endophytic bacteria and alpha-rhizobia in their root nodules.
E. contorsiliquum may establish a nitrogen-fixing symbiosis with Bradyrhizobium lianonginese, and P. parviflora may develop nodules with Bradyrhizobium spp strains that need to be studied further to determine the species. V. caven within the environments sampled may be nodulated by both Ensifer spp and Mesorhizobium strains, suggesting that it is a promiscuous legume species that interacts mainly with broad host range rhizobia.
It is known that nodules are rich in nutrients providing a habitat for bacteria, not only for nitrogen-fixing rhizobia, but for other microorganisms (associated or endophytic) as well, and interestingly, the native legumes studied contained Gammaproteobacteria (Pseudomonas, Dyella, Caulobacter), Actinobacteria (Curtobacterium, Micobacterium) and Firmicutes (Bacillus), that in in-vitro studies showed they can control fungal pathogens or solubilize P. These non-rhizobia endophytes might play a role in nutrition, promoting plant growth and health. In this regard, more work is needed to understand the role of the endophytic bacteria in root nodules of these legumes in combination with rhizobia, as well as to establish their condition as micro-symbiont or endophytic bacteria.
These biological roles of symbiotic and endophytic microorganisms' interactions as well as the knowledge of nodule microbiomes might contribute to have a better understanding of the system, and a better management of natural resources.
Acknowledgments
Virginia Martínez
Alcántara is a teaching researcher of the Agricultural Microbiology chair, School of Agricultural and Forestry Sciences, and Centro de Investigaciones de Fitopatologia (CIDEFI), Universidad Nacional de La Plata (UNLP), Argentina.
Rocío Medina is a fellow of the Consejo Nacional de
Investigaciones Científicas y Técnicas (CONICET), Argentina, and CIDEFI. Juan Gauna is a fellow student of Microbiologia Agrícola Facultad de Ciencias Agrarias y
Forestales. Pedro A. Balatti is
a research scientist at
Comisión de Investigaciones Cientificas de la Provcincia de Buenos Aires, and full professor of the Agricultural Microbiology chair, and
full profesor Facutlad de Ciencias Agrarias y
Forestales, Universidad Nacional de La Plata. This work was supported by a GRANT provided by a Research program of the Secretaría de Políticas Universitarias Ministerio de Eduación Argentina
References
1. Peix A, Ramírez-Bahena MH, Velázquez E, Bedmard EJ. Bacterial associations with Legumes. Crit Rev Plant Sci. 2015;34:17-42.
2. Raven PH, Polhill RM. Advances in Legume Systematics: International Legume Conference; 31 July-4 August 1978; Kew, England. Kew (GB): Royal Botanic Gardens; 1981. 425p.
3. Doyle JJ, Luckow MA. The rest of the iceberg : legume diversity and evolution in a phylogenetic context. Plant Physiol. 2003;131:900-10.
4. Lewis G, Schrire B, MacKinder B, Lock M, editors. Legumes of the World. Kew (GB): Royal Botanic Gardens; 2005. 577p.
5. Franche C, Lindström K, Elmeric C. Nitrogen-fixing bacteria associated with leguminous and non-leguminous plants. Plant Soil. 2009;32:35-59.
6. Sprent JI. Evolving ideas of legume evolution and diversity: a taxonomic perspective on the occurrence of nodulation. New Phytol. 2007;174:11-25.
7. Sprent JI, Ardley JK, James EK. From North to South: a latitudinal look at legume nodulation processes. S Afr J Botany. 2013;89:31-41.
8. Gyaneshwar P, Hirsch AM, Moulin L, Chen WM, Elliott GN, Bontemps C. Legume-Nodulating Betaproteobacteria: diversity, host range, and future prospects. Mol Plant-Microbe Interact. 2011;24(11):1276-88.
9. Compant S, Clément C, Sessitsch A. Plant growth-promoting bacteria in the rhizo-and endosphere of plants: their role, colonization, mechanisms involved and prospects for utilization. Soil Biol Biochem. 2010;42(5):669-78.
10. Glick B. Plant growth-promoting bacteria: mechanisms and applications. Scientifica [Internet]. 2012 [cited 2020 Oct 19];2012:963401. Aavailable from: https://bit.ly/35gCHtm.
11. Laranjo M, Alexandre A, Oliveira S. Legume growth-promoting rhizobia: an overview on the Mesorhizobium genus. Microbiol Res. 2014;169(1):2-17.
12. Frioni L, Dodera R, Malate´s D, Irigoyen I. An assessment of nitrogen fixation capability of leguminous trees in Uruguay. Appl Soil Ecol. 1998;7:271-9.
13. De Faria SM, Diedhiou AG, de Lima HC, Ribeiro RD, Galiana A, Castilho AF, Henriques JC. Evaluating the nodulation status of leguminous species from the Amazonian forest of Brazil. J Exp Bot. 2010;61(11):3119-27.
14. Cordero I, Ruiz-Díeza B, Coba de la Peña T, Balaguerb L, Lucasa MM, Rincóna A, Pueyo JJ. Rhizobial diversity, symbiotic effectiveness and structure of nodules of Vachellia macracantha. Soil Biol Biochem. 2016;96:39-54.
15. Chen WM, Moulin L, Bontemps C, Vandamme P, Béna G, Boivin-Masson C. Legume symbiotic nitrogen fixation by β-proteobacteria is widespread in nature. J Bacteriol. 2003;185(24):7266-72.
16. Chen WM, De Faria SM, Straliotto R, Pitard RM, Simoes-Araujo JL, Chou JH, Sprent JI. Proof that Burkholderia strains form effective symbioses with legumes: a study of novel Mimosa-nodulating strains from South America. Appl Environ Microbiol. 2005;71(11):7461-71.
17. Chen WM, James EK, Coenye T, Chou JH, Barrios E, De Faria SM, Vandamme P. Burkholderia mimosarum sp. nov., isolated from root nodules of Mimosa spp. from Taiwan and South America. Int J Syst Evol Microbiol. 2006;56(8):1847-51.
18. Sprent JI, Ardley J, James EK. Biogeography of nodulated legumes and their nitrogen‐fixing symbionts. New Phytol. 2017;215(1):40-56.
19. De Oliveira ISR, Jesus EDC, Ribeiro TG, da Silva MSR, Tenorio JO, Martins LMV, Faria SM. Mimosa caesalpiniifolia Benth. adapts to rhizobia populations with differential taxonomy and symbiotic effectiveness outside of its location of origin. FEMS Microbiol Ecol [Internet]. 2019 [cited 2020 Oct 19]; 95(8):fiz109. Available from: https://bit.ly/3dJ2KNF.
20. Frioni L, Rodríguez A, Meerhoff M, de los Santos C. Biodiversity in Rhizobia from Native Legume Trees from Uruguay. In: Pedrosa FO, Hungria M, Yates G, Newton WE, editors. Nitrogen fixation: from molecules to crop productivity. Dordrecht: Springer; 2002. p. 293.
21. Langleib M, Beracochea M, Zabaleta M, Battistoni F, Sotelo-Silveira J, Fabiano E, Iriarte A, Platero R. Draft Genome Sequence of Paraburkholderia sp. UYCP14C, a Rhizobium Strain Isolated from Root Nodules of Calliandra parvifolia. Microbiol Resour Announc [Internet]. 2019 [cited 2020 Oct 19]; 8(16):e00173-19. Available from: https://bit.ly/3lTaOhQ.
22. Pereira-Gómez M, Ríos C, Zabaleta M, Lagurara P, Galvalisi U, Iccardi P, Azziz G, Battistoni F, Platero R, Fabiano E. Native legumes of the Farrapos protected area in Uruguay establish selective associations with rhizobia in their natural habitat. Soil Biol Biochem [Internet]. 2020 [cited 2020 Oct 19];140:107854. Avaialble from: https://bit.ly/34bPWMU. Subscription required to view.
23. Milnitsky F, Frioni L, Agius F. Characterization of rhizobia that nodulate native legume trees from Uruguay. Soil Biol Biochem. 1997;29(5-6):989-92.
24. Moreira FM, Haukka K, Young JP. Biodiversity of rhizobia isolated from a wide range of forest legumes in Brazil. Mol Ecol. 1998;7(7):889-95.
25. Paulitsch F, Dall'Agnol RF, Delamuta JRM, Ribeiro RA, da Silva Batista JS, Hungria M. Paraburkholderia guartelaensis sp. nov., a nitrogen-fixing species isolated from nodules of Mimosa gymnas in an ecotone considered as a hotspot of biodiversity in Brazil. Arch Microbiol. 2019;201(10):1435-46.
26. Platero R, James EK, Rios C, Iriarte A, Sandes L, Zabaleta M, Battistoni F, Fabianoa E. Novel Cupriavidus Strains Isolated from Root Nodules of Native Uruguayan Mimosa Species. Appl Environ Microbiol. 2016;82(11):3150-64.
27. Silva VC, Alves PAC, Rhem MFK, Dos Santos JMF, James EK, Gross E. Brazilian species of Calliandra Benth. (tribe Ingeae) are nodulated by diverse strains of Paraburkholderia. Syst Appl Microbiol. 2018;41(3):241-50.
28. Taulé C, Zabaleta M, Mareque C, Platero R, Sanjurjo L, Sicardi M, Frioni L, Battistoni F, Fabiano E. New betaproteobacterial Rhizobium strains able to efficiently nodulate Parapiptadenia rigida (Benth.) Brenan. Appl Environ Microbiol. 2012;78(6):1692-700.
29. Frioni L, Malatés D, Irigoyen I, Dodera R. Promiscuity for nodulation and effectivity in the N2-fixing legume tree Acacia caven in Uruguay. Appl Soil Ecol. 1998;7(3):239-44.
30. De Faria SM, Franco AA, de Jesús RM, Menandro DS, Baitello JB, Mucci ESF. New nodulating legume trees from south-east brazil. New Phytol. 1984;98:317-28.
31. Ferreira Fontenele AC, Menezes Aragão W, de Albuquerque Rangel JH, Aragão Almeida S. Leguminosas tropicais: Desmanthus virgatus (L.) Willd., uma forrageira promisora. Rev bras Agrocienc. 2009;15(1-4):121-3.
32. Burkart R, Bárbaro NO, Sánchez RO, Gómez DA. Eco-regiones de la Argentina. Buenos Aires: Administración de Parques Nacionales; 1999. 42p.
33. Vincent JM. A manual for the practical study of root-nodule bacteria. Oxford: Blackwell Scientific; 1970. 164p.
34. Brown VI, Lowbury RJL. Use of an improved cetrimide agar medium and other culture methods for pseudomonas aeruginosa. J Clin Pathol. 1965;18:752-6.
35. Standard Methods for the Examination of Dairy Products. 14th ed. Washington (DC): APHA; 1978. 416p.
36. Velazquez E, Igual JM, Willems A, Fernández MP, Munoz E, Mateos PF. Mesorhizobium chacoense sp. nov., a novel species that nodulates Prosopis alba in the Chaco Arido region (Argentina). Int J Syst Evol Microbiol. 2001;51:1011-21.
37. Colco R. Gram Staining. Curr Protoc Microbiol [Internet]. 2005 [cited 2020 Oct 19];00(1). Available from: https://bit.ly/34byLLd. Subscription required to view.
38. Nautiyal CS. An efficient microbiological growth medium for screening phosphate solubilizing microorganisms. FEMS Microbiol Lett. 1999;170(1):265-70.
39. Franco MEE, Troncozo MI, López SMY, Lucentini G, Medina R, Saparrat MCN, Balatti PA. A survey on tomato leaf grey spot in the two main production areas of Argentina led to the isolation of Stemphylium lycopersici representatives which were genetically diverse and differed in their virulence. Eur J Plant Pathol. 2017;149(4):983-1000.
40. Malbrán I, Mourelos CA, Balatti PA, Lori GA. Aggressiveness and genetic variability of Fusarium graminearum populations from the main wheat production area of Argentina. J King Saud Univ Sci. 2019;31(4):1215-9.
41. Atlas RM. Handbook of Microbiological Media. 2nd ed. Boca Ratón: CRC Press; 1997. 1712p.
42. UTHSCSA ImageTool [Internet]. Version 3.0. San Antonio (TX): UTHSCSA; 1995-2002 [cited 2020 Oct 20]. Available from: https://bit.ly/3dK5tGJ.
43. Plazas-Jiménez OD, Coy-Barrera E. Antifungal activity of ultrasound-assisted acetone extract and alkylresorcinol-enriched fractions from Hordeum vulgare L. against Fusarium oxysporum. Cul Trop. 2016;37(4):173-81.
44. Di Rienzo JA, Casanoves F, Balzarini M, Laura G, Margot T, Robledo C. InfoStat [Internet]. Version 07-10-2018. Córdoba: Universidad Nacional de Córdoba, Facultad de Ciencias Agropecuarias; 2014 [cited 2020 Oct 19]. Available from: https://bit.ly/3dDvIyu.
45. López S, Pastorino G, Franco M, Medina R, Lucentini C, Saparrat M, Balatti P. Microbial endophytes that live within the seeds of two tomato hybrids cultivated in Argentina. Agron. 2018;8(8):136.
46. Medina R, López SM, Franco ME, Rollan C, Ronco BL, Saparrat MC, Balatti PA. A Survey on occurrence of Cladosporium fulvum identifies race 0 and race 2 in tomato-growing areas of Argentina. Plant diseas. 2015;99(12):1732-7.
47. López S, Balatti PA. Closely related strains of Bradyrhizobium contained in commercial inoculates of soybean are identified by a set of PCR reactions. J Genet Eng Biotechnol [Internet]. 2011 [cited 2020 Oct 20]; 2011: 8p. Available from: https://bit.ly/35jp7W3.
48. Hammer Ø, Harper DAT, Ryan PD. PAST: Paleontological statistics software package for education and data analysis. Palaeontologia Electronica. 2001;4(1):9p.
49. Reysenbach AL, Wickham GS, Pace NR. Phylogenetic analysis of the hyperthermophilic pink filament community in Octopus Spring, Yellowstone National Park. Appl Environ Microbiol. 1994;60(6):2113-9.
50. Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor (NY): Cold Spring Harbor Laboratory Press; 1989. 3v.
51. Geneious [Internet]. Version R9.2.5. 2019 [cited 2020 Oct 20]. Available from: https://bit.ly/34iP8pa.
52. Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22(22):4673-80.
53. Tamura K, Nei M, Kumar S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA. 2004;101(30):11030-5.
54. Arturi MF, Juárez MC. Composición de las comunidades arbóreas de la Isla Martín García en relación a un gradiente ambiental. Ecol Austral. 1997;7:65-72.
55. Barotto AJ, Cellini JM. Quantitative analysis of the floristic composition of the Martín García Island. In: XVIII jornadas técnicas forestales y ambientales; 2019 Oct 17-18; Eldorado, Misiones, Argentina. Misiones: Facultad de Ciencias Forestales; 2019. p. 243-5.
56. Andrews M, Andrews ME. Specificity in Legume-Rhizobia Symbioses. Int J Mol Sci. 2017;18(4):705.
57. Vinuesa P, Silva C, Lorite MJ, Izaguirre-Mayoral ML, Bedmar EJ, Martínez-Romero E. Molecular systematics of rhizobia based on maximum likelihood and Bayesian phylogenies inferred from rrs, atpD, recA and nifH sequences, and their use in the classification of Sesbania microsymbionts from Venezuelan wetlands. Syst Appl Microbiol. 2005;28:702-16.
58. Remigi P, Zhu J, Young JPW, Masson-Boivin C. Symbiosis within symbiosis: evolving nitrogen-fixing legume symbionts. Trends Microbiol. 2016;24:63-75.
59. Benhizia Y, Benhizia H, Benguedouar A, Muresu R, Giacomini A, Squartini A. Gamma proteobacteria can nodulate legumes of the genus Hedysarum. Syst Appl Microbiol. 2004;27(4):462-8.
60. Muresu R, Polone E, Sulas L, Baldan B, Tondello A, Delogu G, Cappuccinelli P, Alberghini S, Benhizia Y, Benhizia H, Benguedouar A, Mori B, Calamassi R, Dazzo FB, Squartini A. Coexistence of predominantly non culturable rhizobia with diverse, endophytic bacterial taxa within nodules of wild legumes. FEMS Microbiol Ecol. 2008;63:383-400.
61. Shiraishi A, Matsushita N, Hougetsu T. Nodulation in black locust by the Gammaproteobacteria Pseudomonas sp. and the Betaproteobacteria Burkholderia sp. Syst Appl Microbiol. 2010;33(5):269-74.
62. Huang B, Lv C, Zhao Y, Huang R. A Novel Strain D5 Isolated from Acacia confusa. PLoS ONE [Internet]. 2012 [cited 2020 Oct 20]; 7(11):e49236. Available from: https://bit.ly/31moUAf.
63. Ampomah OY, Huss-Danell K. Genetic diversity of root nodule bacteria nodulating Lotus corniculatus and Anthyllis vulneraria in Sweden. Syst Appl Microbiol. 2011;34:267-75.
64. Saïdi S, Chebil S, Gtari M, Mhamdi R. Characterization of root-nodule bacteria isolated from Vicia faba and selection of plant growth promoting isolates. World J Microbiol Biotechnol. 2013;29(6):1099-106.
65. Truchet G, Barker DG, Camut S, Billy F, Vasse J, Huguet T. Alfalfa nodulation in the absence of Rhizobium. Mol Gen Genet. 1989;219:65-8.
66. Zgadzaj R, James EK, Kelly S, Kawaharada Y, de Jonge N, Jensen DB, Madsen LH, Radutoiu S. A Legume Genetic Framework Controls Infection of Nodules by Symbiotic and Endophytic Bacteria. PLoS Genet [Internet]. 2015 [cited 2020 Oct 20]; 11(6):e1005280. Available from: https://bit.ly/3dHX9at.
67. Binde DR, Menna P, Bangel EV, Barcellos FG, Hungria M. Rep-PCR fingerprinting and taxonomy based on the sequencing of the 16S rRNA gene of 54 elite commercial rhizobial strains. Appl Microbiol Biotechnol. 2009;83:897-908.
68. Laranjo M, Young JPW, Oliveira S. Multilocus sequence analysis reveals multiple symbiovars within Mesorhizobium species. Syst Appl Microbiol. 2012;35:359-67.
69. Alexandre A, Brígido C, Laranjo M, Rodrigues S, Oliveira S. Survey of Chickpea Rhizobia Diversity in Portugal Reveals the Predominance of species distinct from Mesorhizobium ciceri and Mesorhizobium mediterraneum. Microbial Ecol. 2009;58:930-41.
70. Beukes CW, Boshoff FS, Phalane FL, Hassen AI, Le Roux MM, Stȩpkowski T, Venter SN, Steenkamp ET. Both Alpha- and Beta-Rhizobia occupy the Root Nodules of Vachellia karroo in South Africa. Front Microbiol [Internet]. 2019 [cited 2020 Oct 20];10:1195. Available from: https://bit.ly/3j9FBFm.
71. Pueppke SG, Broughton WJ. Rhizobium sp. strain NGR234 and R. fredii USDA257 share exceptionally broad, nested host ranges. Mol Plant Microbe Interact. 1999;12(4):293-318.
72. Choudhary S, Tak N, Bissa G, Chouhan B, Choudhary P, Sprent JI, James EK, Gehlot HS. The widely distributed legume tree Vachellia (Acacia) nilotica subsp. indica is nodulated by genetically diverse Ensifer strains in India. Symbiosis. 2020;80:15-31.
73. Sankhla IS, Tak N, Meghwal RR, Choudhary S, Tak A, Rathi S, Sprent JI, James EK, Gehlot HS. Molecular characterization of nitrogen fixing microsymbionts from root nodules of Vachellia (Acacia) jacquemontii, a native legume from the Thar Desert of India. Plant Soil. 2017;410:21-40.
74. Parte AC. LPSN - List of Prokaryotic names with Standing in Nomenclature (bacterio.net), 20 years on. Int J Syst Evol Microbiol [Internet]. 2018 [cited 2020 Oct 20];68:1825-9. Available from: https://bit.ly/34cwoYD.
75. Wang ET, van Berkum P, Sui XH, Beyene D, Chen WX, Martinez-Romero E. Diversity of rhizobia associated with Amorpha fruticosa isolated from Chinese soils and description of Mesorhizobium amorphae sp. nov. Int J Syst Bacteriol. 1999;49:51-65.
76. Zhao CT, Wang ET, Zhang YM, Chen WF, Sui XH, Chen WX, Liu HC, Zhang XX. Mesorhizobium silamurunense sp. nov., isolated from root nodules of Astragalus species. Int J Syst Evol Microbiol. 2012;62:2180-6.
77. Zhu YJ, Kun J, Chen YL, Wang SK, Sui XH, Kang LH. Mesorhizobium acaciae sp. nov., isolated from root nodules of Acacia melanoxylon R. Br. Int J Syst Evol Microbiol. 2015;65:3558-63.
Supplementary material
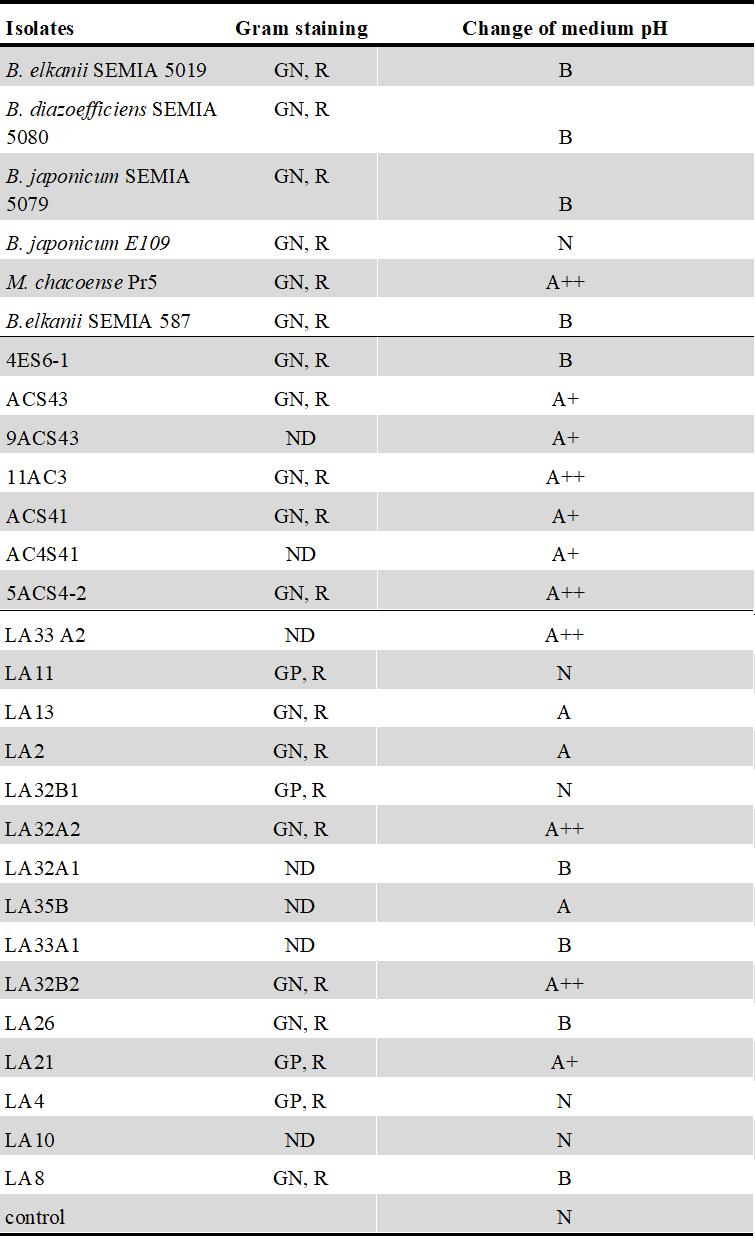
Table S1
Microbiological characterization of 22
isolates and 5 reference strains used in this study.
Control treatment consisted in culture medium without bacteria. GN, Gram-negative; GP, Gram-positive; R, rods; A++, strong acid;
A+, moderately acid; A, slightly acid; N, neutral; B, basic; ND, No Determined
Author notes
vmalcan@gmail.com
Additional information
Author
contribution statement: Virginia Martínez Alcántara
and Rocío Medina wrote the manuscript and, in
collaboration with Juan Gauna, made the data
analysis, lab and field work. Pedro Balatti
collaborated in planning the experiments, the analysis and writing the
manuscript.
Editor: The
following editors approved this article.
Alicia Castillo
Instituto Nacional de Investigación Agropecuaria
(INIA), Canelones, Uruguay.
ORCID: https://orcid.org/0000-0001-7381-3887 // Paul Chavarriaga
Centro Internacional de Agricultura Tropical (CIAT),
Cali, Colombia.
ORCID: https://orcid.org/0000-0001-7579-3250
Alternative link
http://agrocienciauruguay.uy/ojs/index.php/agrociencia/article/view/411/416 (pdf)











